Customize Library Upload Library
Resources: Download Assessment Tool SWATH Libraries Publications
About SWATH MS
Related: PeptideAtlas SRMAtlas PASSEL
LOG IN

DIALib-QC v1.2 is a tool that allows users to analyze DIA libraries in a variety of formats including PeakView, OpenSWATH, Spectronaut and Predicted ion libraries (from Prosit- Spectronaut compatible generic format in csv) based on a library characteristics.
These include charge, length, retention time, m/z, complexity, etc. In addition to assessment, users can correct the ion library for any mass error and/or remove conflict assays. The supported libraries can be assessed online at SWATHAtlas for moderately-sized libraries (up to 200 MB), otherwise a downloaded version can be run locally on most Linux systems.
The DIALib-QC v1.2 tool is written in the perl programming language, so perl installation is a pre-requisite on your Linux system. There is one non-core module dependency, Statistics::Regression. The tool will work without it, but you will lose the test comparing RT values of +2 and +3 ions of the same modified peptide, a metric of the retention time consistency of the ion library being analyzed. Instructions for perl module installation are at available at the README file below. Once you've downloaded the script, you can execute the file directly, or as an argument to perl.
Files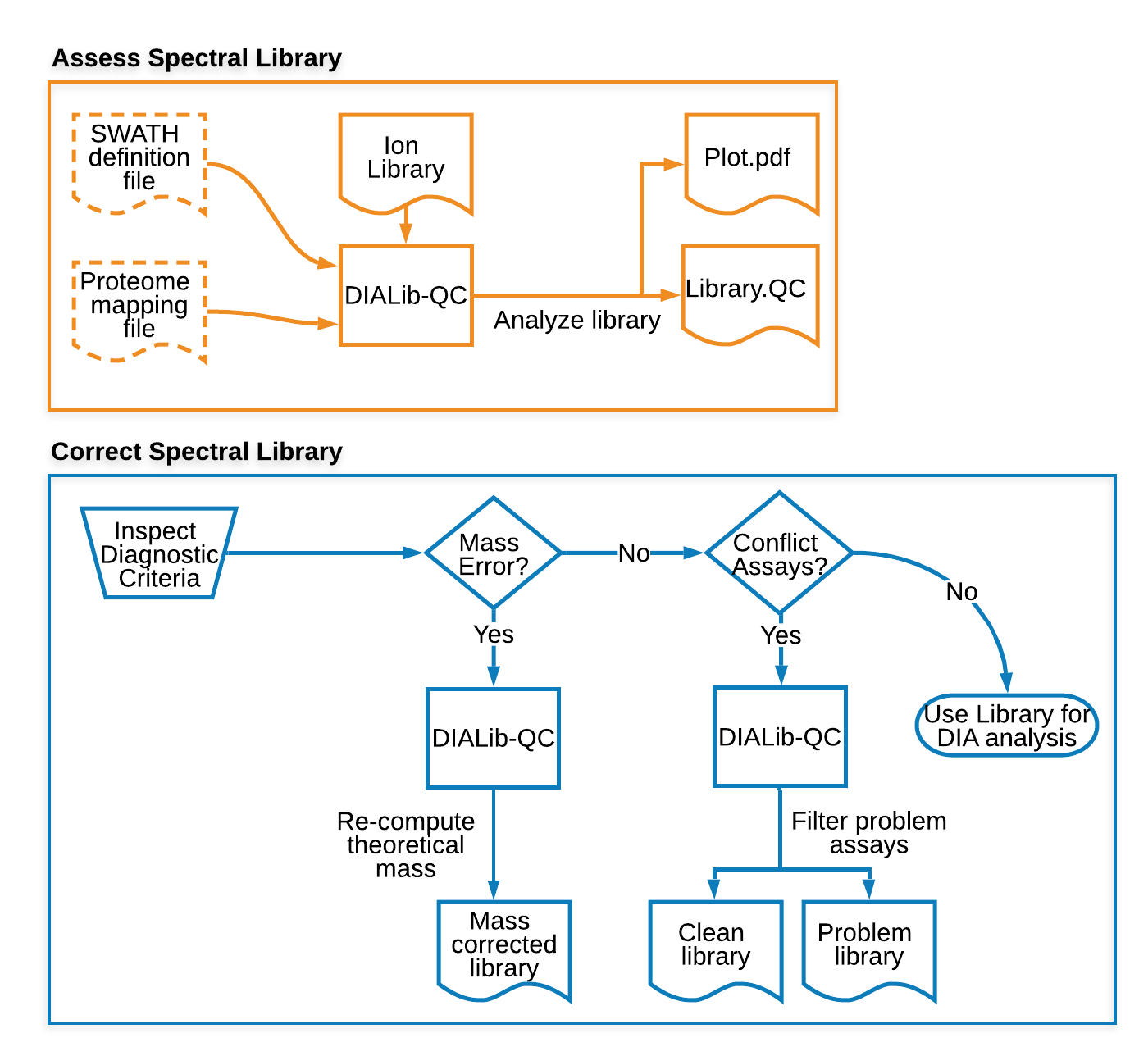
Below is the DIALib-QC v1.2 usage protocol or workflow to assess and repair a spectral ion library. The current version of the DIALib-QC supports the following list of modifications (modification annotation patterns).
Here we present a spectral ion library (K562_q3bad_PV) referring from the Midha et. al. paper published in Nature Communications (2020).
All the below mentioned steps work well with PeakView, Spectronaut, OpenSWATH and In-silico predicted library (from Prosit- generic csv (Spectronaut compatible)) formats. We are providing examples of two spectral ion libraries format (PeakView and In-silico ion library), SWATH_definitions, peptide mapping file and expected result files via the links below.